Research Projects
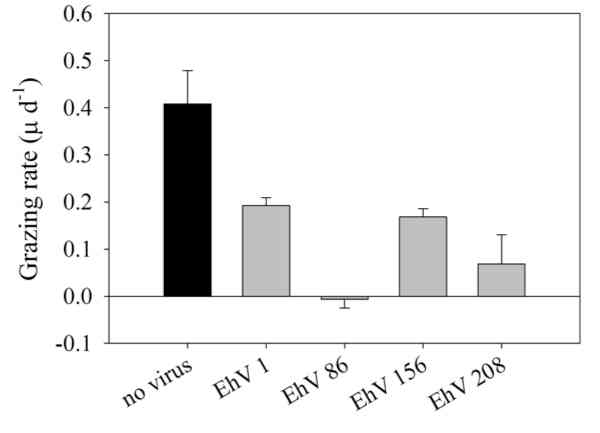
Quantifying competing loss rates of viral lysis and microzooplankton grazing on Emiliania huxleyi mortality
NSF Biological Oceanography
Matt Johnson1, Elizabeth Harvey1,2, and Kay Bidle3
1Woods Hole Oceanographic Institution; 2Skidaway Institute of Oceanography, University of Georgia; 3Institute of Marine and Coastal Sciences, Rutgers University
The coccolithophore, Emiliania huxleyi plays a large role in global carbon cycling, as their calcite coccoliths account for a third of marine CaCO3 production. Like all phytoplankton, the abundance and distribution of E. huxleyi is driven by processes that stimulate growth and expedite mortality.
Read more
While the two main mechanisms of mortality in E. huxleyipopulations are microzooplankton grazing and viral lysis, the latter has receive much greater attention in recent years. Ultimately these processes may have contrasting influence on the structure and function of the marine food web and biogeochemical cycles. Grazing channels phytoplankton biomass to higher trophic levels, whereas viral lysis stimulates vertical particle export flux. The investigators hypothesize that microzooplankton grazing is the primary cause of E. huxleyi mortality, but that grazing-induced mortality decreases over the course of an E. huxleyi bloom, whereas loss to viral lysis becomes more important. To test this hypothesis, we will conduct controlled laboratory experiments where the influence of E. huxleyi viral infection on microzooplankton grazing rates is examined, as well as the role of grazing in enhancing viral infection dynamics. After mechanistically characterizing these interactions, the contribution of both processes to E. huxleyi mortality and particle export flux will be quantified in both laboratory and field mesocosm experiments.
Phytoplankton are one of the central components in controlling carbon cycling in the marine environment, thus, shifts in their population distribution and abundance can have a large impact on biogeochemical cycling and global climate change. As phytoplankton population dynamics are driven by the relative rates of cell growth and loss, interactions with other members of the planktonic community can directly influence both of these rate processes. E. huxleyi is a model organism for examining how mortality is partitioned between microzooplankton grazing and viral lysis, as it forms massive annual blooms that contribute greatly to global carbon fluxes, and it also has a well-established history with a lytic virus. Deciphering the nature and relative importance of these cell-cell-virus interactions will help to illuminate mechanisms that drive bulk measurements of phytoplankton loss and is necessary to seed models for the accurate prediction of phytoplankton population dynamics and ultimately, biogeochemical cycling and shifts in global climate.
Exploring the physiological and ecological basis of mixotrophy in marine food webs
NSF Biological Oceanography
Matt Johnson, Woods Hole Oceanographic Institution
This project aims to determine the cellular and environmental factors that lead to feeding by marine phytoplankton, i.e. mixotrophy, and how the contrasting metabolisms of heterotrophy and photosynthesis are integrated within a cell.
Read more
Using microscopy, physiology, proteomic, and metabolomic methods, we will test hypotheses about the causes and consequences of mixotrophy. The major objectives of this proposal are to determine 1) environmental controls for inducing mixotrophy, 2) the role of prey quality on predator selection, 3) cellular and molecular controls of mixotrophy, and 4) nutrient assimilation and integrated metabolism. Using these various research approaches, we will produce a comprehensive view of several mixotrophs and provide new insights into cellular, ecological, and evolutionary aspects of mixotrophy.
Mixotrophy refers to species that combine some level of phagotrophy (feeding) and phototrophy (photosynthesis), and represents a diverse array of ecological interactions and cellular and metabolic adaptations. The proposed research focuses on phytoplankton that feed, and uses representative dinoflagellate and chrysophyte species. While often perceived as an exception to the norm, mixotrophy is commonplace in marine food webs. Furthermore, phagotrophy is an ancestral trait of all eukaryotes, and is responsible for the high diversity of eukaryotic phytoplankton, through endosymbiotic chloroplast acquisitions. Mixotrophy affords phytoplankton greater ecological fitness during periods of low or limiting nutrients while stabilizing food webs, and there is mounting evidence that it is ubiquitous in low nutrient ecosystems such as ocean gyres. Many mixotrophs have a low chlorophyll: carbon ratio, which tends to make them poor phototrophic competitors. In turn, feeding allows them to achieve their maximum growth while in some cases also eliminates their competitors. Other mixotrophs are strong phototrophic competitors, and only feed when severely nutrient limited. While it is known that mixotrophs are ecologically important, we have a limited understanding of the cellular mechanisms regulating mixotrophy. Such information is key for predicting the role of mixotrophy through the development of ecosystem and mechanistic models. The proposed research will help to clarify when and why mixotrophs feed, and how heterotrophy and photoautotrophy are integrated within phytoplankton cells. Key to this research is the poorly understood relationship between prey and predator nutrient limitation (e.g. stoichiometry), and how these factors affect feeding dynamics. Further, the proposed research will distinguished between two major modes of mixotrophy, i.e. inducible and persistent, which can be distinguished by a cell’s dependency upon heterotrophy, its propensity to feed, and ultimately by its ecological niche or function. Results from this project will improve our understanding of the physiological and ecological role of mixotrophy in marine phytoplankton, and provide much needed molecular markers for studying this process in both the laboratory and field.
Cellular and metabolic integration of symbiotic organelles in the ciliate Mesodinium rubrum NSF Integrative Organismal Systems
Matt Johnson1 and and Erica Lasek-Nesselquist2
1Woods Hole Oceanographic Institution , 2University of Scranton
The proposed research will determine how the ciliate, Mesodinium rubrum, is capable of maintaining and replicating stolen cryptophyte algal organelles. Central to this phenomenon, is the ciliate’s ability to steal and temporarily maintain an active nucleus.
Read more
The project will utilize traditional and next generation omics research approaches, to provide an unprecedented perspective of a model evolutionary “chimera”. The objectives of this proposal are to understand 1) how cryptophyte organelles are organized in M. rubrum and their fate, 2) how M. rubrum orchestrates gene and protein expression of symbiotic organelles, and 3) how metabolic pathways in M. rubrum are integrated and adapted to photosynthesis. By combining a variety of microscopic and cell-targeted techniques with transcriptomics, proteomics, and metabolomics, this project will provide a comprehensive view of the ciliate’s integrated metabolism and how it becomes perturbed when the cryptophyte nucleus is lost and under stress treatments.
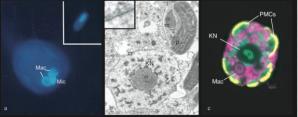
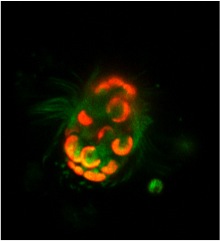
The temporary acquisition of phototrophy, which includes the processes of algal endosymbiosis or stealing their chloroplasts (kleptoplasty), is widespread in aquatic ecosystems and distributed broadly across the eukaryotic tree of life. These phenomena are important adaptations that allow organisms to exploit phototrophy, thereby increasing their biochemical potential and growth efficiency. However, it is also a potential evolutionary path to the stable acquisition of a chloroplast (e.g. secondary endosymbiosis). M. rubrum is a globally distributed mixotrophic ciliate found in marine coastal environments, and is capable of forming non-toxic and highly productive red tides. The ciliate was long considered an enigma due to the conspicuous presence of symbiotic chloroplasts, mitochondria, and other organelles of cryptophyte origin. We now know that these organelles are ultimately stolen from free-living cells. Like endosymbiosis, organelle-retention increases fitness of a host by providing excess energy at little cost, but typically requires frequent replacement of stolen chloroplasts. However, M. rubrum has the unique capacity to control and divide these organelles in a symbiotic state. This is due to the ciliate’s ability to steal and temporarily maintain a functional cryptophyte nucleus- a process termed karyoklepty. The sequestered cryptophyte nucleus in M. rubrum cannot divide, thus the ciliate needs to reacquire it by feeding. When present, the nucleus is transcriptionally active and the ciliate can apparently facilitate protein transport to the chloroplasts.
Ultimately, this research will help to determine how a non-photosynthetic organism has adapted to obligate phototrophy- a process that is recurrent and profoundly important in eukaryotic evolution. At its heart this proposal seeks to illuminate these evolutionary processes and will have broad implications for understanding diverse endosymbiotic interactions. Results from this project will provide an integrative picture of an acquired phototroph that appears to be at an evolutionary crossroads- adapted to phototrophy but lacking full control of the genetic machinery needed to power its metabolism. M. rubrum is an ideal model for understanding stable plastid acquisitions, due to its practice of karyoklepty and its poorly understood protein-targeting system. Finally, this research will provide an unprecedented view of an ecologically important organism, which has long been viewed as a functional enigma.
Infochemical control of microbial interactions and nutrient cycling in the North Atlantic
Gordon and Betty More Foundation
Benjamin Van Mooy1*, Kay Bidle2, Mathew Johnson1, Tracy Mincer1, Assaf Vardi3.
1 Woods Hole Oceanographic Institution, Woods Hole, MA.
2Institute of Marine and Coastal Sciences, Rutgers University, New Brunswick, NJ.
3Department of Plant Sciences, Weizmann Institute of Science, Rehovot, Israel.
* Lead-PI
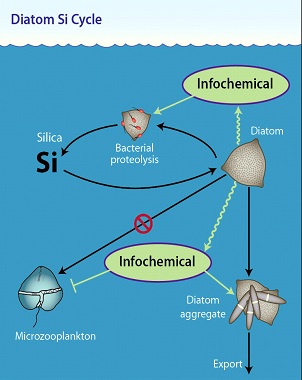
Our overarching scientific objective is to understand the role of infochemical signaling on the flow of key nutrients during plankton blooms in the North Atlantic. Building upon our prior work, our project will target a diverse array of cornerstone infochemicals and assess how they critically regulate microbial interactions, ecosystem dynamics and nutrient fluxes.
Read more
We have identified enigmatic biogeochemical pathways that consistently defy explanation using canonical bottom-up or top-down ecosystem controls, and instead, propose that they are under ‘infochemical control’. For each pathway we propose a number of testable hypotheses, which will be explored using a threefold approach: first, we will analytically ‘listen’ to infochemical conversations amongst key groups of microbial plankton involved in these pathways; second, we will molecularly ‘interrogate’ these groups in field- and lab-based experiments designed to perturb both environmental conditions and/or signaling pathways to reveal how infochemicals exert controls on cellular metabolism, growth, interactions and mortality at a molecular level; and third, we will biogeochemically ‘implicate’ infochemical signaling as critical controls on biogeochemical fluxes of carbon and nutrients in the sea. We will apply these approaches using novel chemical, genetic, and molecular tools under in situ conditions during NSF-funded cruises.
Completed Projects
Trophodynamics of Mesodinium rubrum and cryptophyte algae
NSF Biological Oceanography
Matt Johnson1 and Diane K. Stoecker2
1Woods Hole Oceanographic Institution; 2Horn Point Lab, University of Maryland, Center for Environmental Science
Mesodinium rubrum (= Myrionecta rubra) is a non-toxic red-tide forming ciliate in coastal and estuarine waters of the world. M. rubrum possesses symbiotic organelles derived from cryptophyte algae, a ubiquitous group of algae in aquatic ecosystems.
Read more
Because ciliates are usually considered “protozoa”- unicellular eukaryotes that have animal rather than plant-like qualities- M. rubrum was long considered an enigma. Studies of phytoplankton ecology have frequently overlooked M. rubrum, instead considering it part of the microzooplankton. In the last decade great progress has been made in understanding how the ciliate functions, largely due to the establishment of cultures. Recent studies have shown that one strain of M. rubrum steal organelles (chloroplasts, mitochondria, and a nucleus) from cryptophyte prey, while another possesses stable (permanent) cryptophyte organelles. All strains of the ciliate, however, must feed on cryptophyte algae, either for acquiring organelles or growth factors. While we now have a greater understanding of the physiology of M. rubrum, almost nothing is known regarding its ecological interactions with cryptophyte algae, bacteria, and potential predators in aquatic ecosystems. Cryptophyte algae are one of the major phytoplankton groups in Chesapeake Bay, contributing to primary production and acting as a major food item for a variety of organisms. M. rubrum is widespread and seasonally abundant in Chesapeake Bay and its tributaries, and can reach red-tide concentrations in the spring and fall. Little is known regarding the genetic diversity of cryptophyte or M. rubrum populations in Chesapeake Bay or other ecosystems.
The objectives of the proposed research are to determine the physical and chemical factors and biological interactions that regulate production of M. rubrum and cryptophytes, and to characterize the range and seasonal patterns of genetic diversity of these organisms in Chesapeake Bay. Another goal is to determine the role of cryptophyte genetic diversity in the production of Mesodinium ciliates. Using oceanographic, physiological, and molecular approaches, we propose to execute the first large scale ecological studies of this unique and ecologically important ciliate, in order to understand its role in marine microbial food webs and to better predict its abundance and distribution.
The proposed research is important for two reasons, 1) M. rubrum can be a dominant member of the plankton in nearly all coastal ecosystems, yet its trophic role remains enigmatic and 2) cryptophyte algae play a pivotal role in the ecology of most estuarine and coastal ecosystems, yet they remain a “black box” of poorly characterized flagellates. Despite numerous reports of M. rubrum red tides, we still have a rudimentary understanding of its mixotrophic role, as both alga and grazer, and how these “strategies” are balanced in natural food webs. The production of M. rubrum, like many other mixotrophs in Chesapeake Bay and other coastal ecosystems, is linked to cryptophyte algal production. However, we are currently unable to predict ecosystem dynamics of M. rubrum, due largely to our lack of knowledge of their cryptophyte prey selection, the spatial and temporal dynamics of cryptophyte production, and the effects of grazing pressure on bloom formation and termination. The Chesapeake Bay is an ideal location to perform this research because both M. rubrum and cryptophytes are extremely abundant, their productivity is seasonally predictable in its timing and location, and sampling sites are easily accessible.
Understanding how Global Warming will select for Zooxanthellae Phenotypes
NSF Biological Oceanography
Paul Falkowski1 and Matt Johnson2
1Institute for Marine and Coastal Sciences; 2Woods Hole Oceanographic Institution
Symbiodinium (= zooxanthellae) is a cosmopolitan dinoflagellate genus of tropical and subtropical latitudes, and an integral member of coral reef ecosystems.
Read more
As the obligate endosymbionts of corals and a variety of other reef organisms, Symbiodinium is the main contributor to coral reef primary production by fueling their host’s metabolism and growth. In this delicate symbiosis the symbiont functions as an autotroph, releasing most of its photosynthetic products as food for their host, while the corals provide protection, a stable environment, and metabolic waste that the algae use to grow. The fragility of this relationship has become increasingly apparent in recent years due to rising ocean temperatures. Coral “bleaching” events, a process by which corals release their symbionts in response to stress, are becoming a global environmental crisis. During this process, Symbiodinium becomes stressed, releasing reactive oxygen species and losing photosynthetic abilities. Prolonged exposure causes irreversible damage and expulsion of zooxanthellae from their hosts. Persistent and recurrent bleaching events can have catastrophic consequences to coral reef health, resulting in the death of coral colonies and a collapse of trophic structure. However, some corals appear to withstand these events unharmed, and laboratory research has shown that some Symbiodinium are tolerant of heat stress. While recent work has revealed a vast genetic diversity of Symbiodinium in coral reef ecosystems, how the diversity is related to stress tolerance is poorly understood.
The objectives of the proposed research are to describe the relationship of Symbiodinium genetic diversity to stress phenotypes in both free-living and symbiotic strains, and to determine how some strains tolerate high temperature stress and resist bleaching in their hosts. Another goal is to find stress-sensitive genotype markers that can be used to evaluate coral reef health and susceptibility to heat stress. Using a combination of biophysical, molecular biological and biochemical assays, the PIs propose to sample the largest number of Symbiodinium strains to date, to experimentally determine how genotype is related to phenotype diversity.
The proposed research is critical for two reasons: 1) Symbiodinium is essential for the health of global coral reef ecosystems and understanding how genetic diversity leads to stress sensitive and tolerant phenotypes will provide a key to understanding which strains are likely to be selected in the coming centuries as a result of bleaching, and 2) very little is known of the relationship between genotype and phenotype diversity in Symbiodinium and most dinoflagellates in general. The first issue is important from a standpoint of conservation biology and reef management. The second provides fundamental understanding of the ecology and evolution of Symbiodinium. Moreover, while Symbiodinium has the smallest genome (based on DNA content) of any non-parasitic dinoflagellates, very little is known regarding gene content, regulation, and expression. Thus, this work will also provide key insights concerning dinoflagellate genome evolution and transcriptional regulation.