Current projects
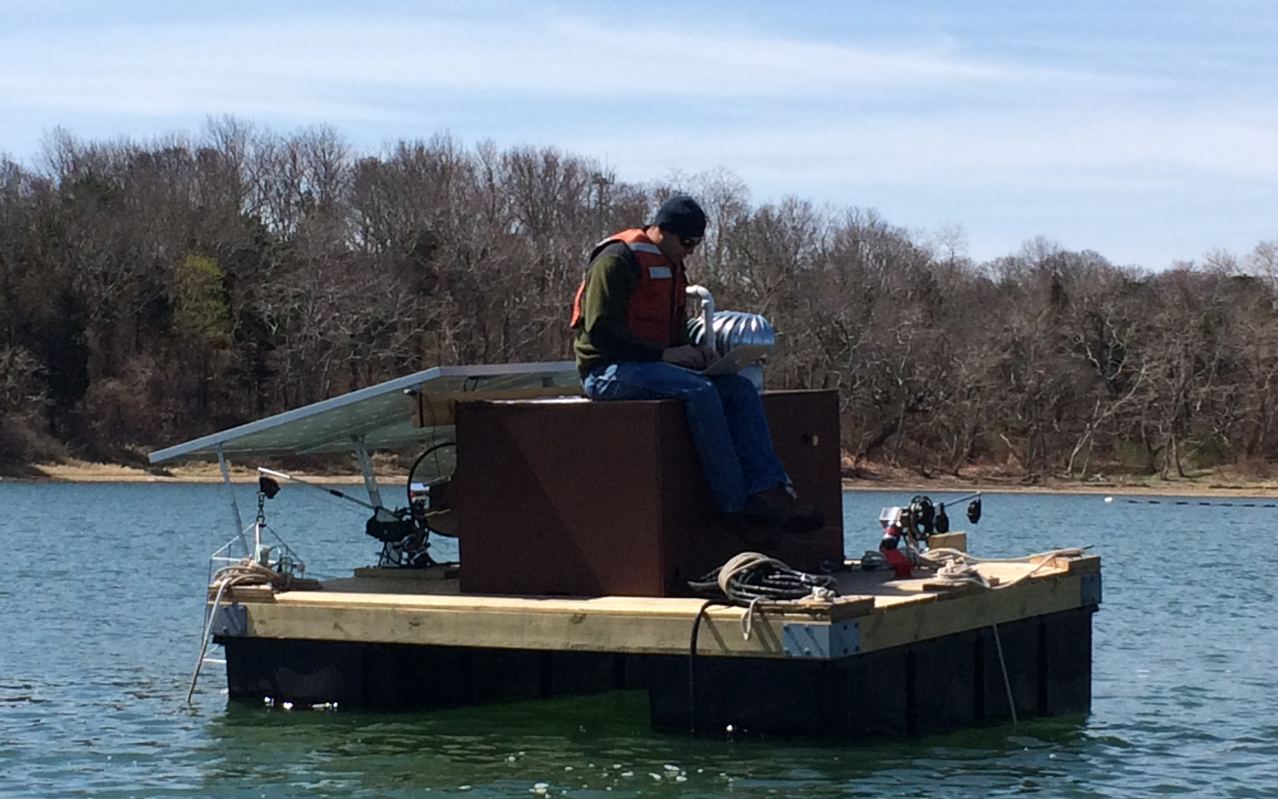

Woods Hole Center for Oceans and Human Health
We are deploying in situ sensors and conducting novel experiments on benthic cysts to characterize the plasticity and temperature dependence of key physiological rates that drive the development and decline of Alexandrium catenella and Pseudo-nitzschia spp. HABs. Check out a neat video about our work here.
HABON-NE, An adaptive observing network for real-time, in situ HAB monitoring and data sharing across New England
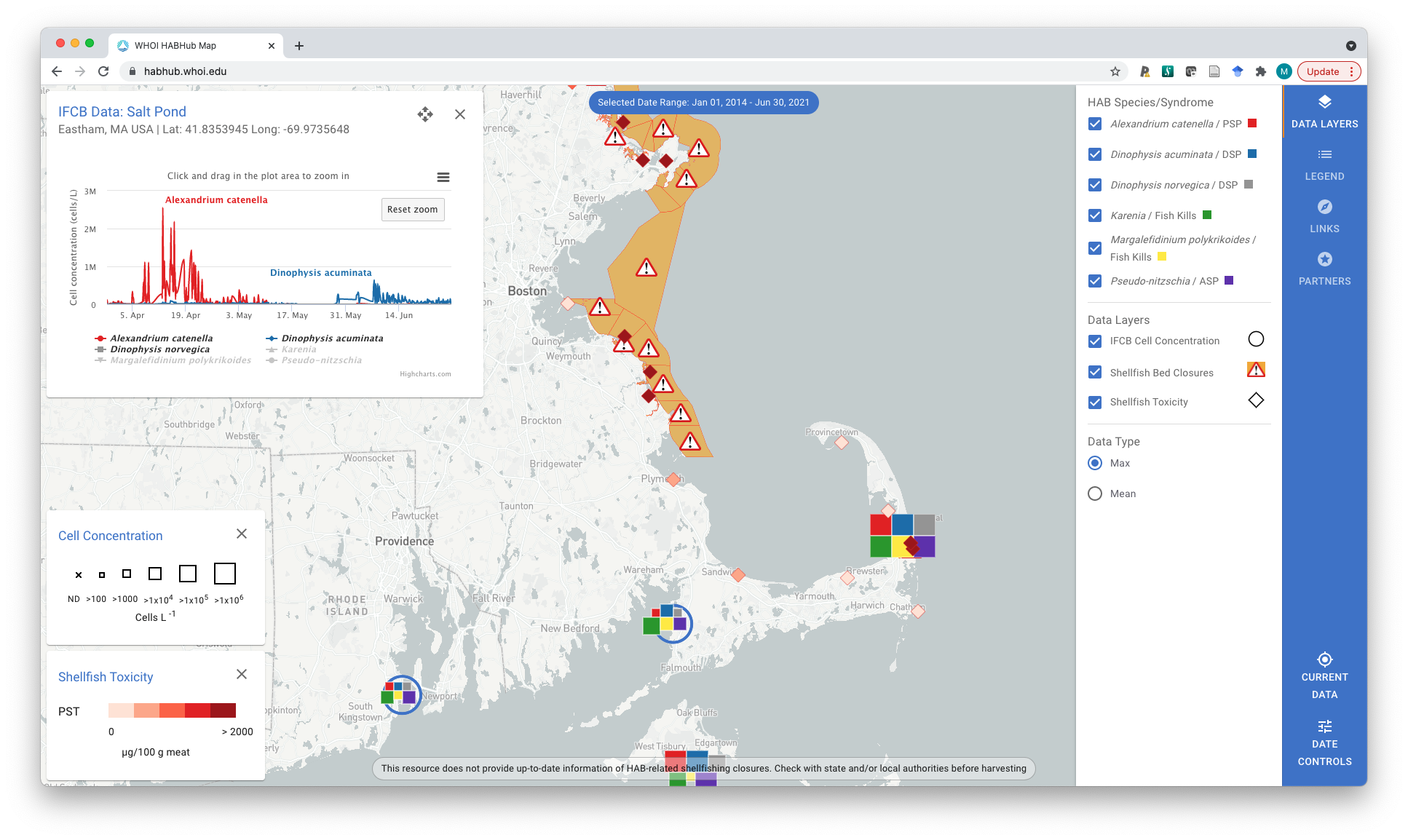
The dinoflagellate Alexandrium catenella, a species that causes paralytic shellfish poisoning (PSP), has long been the dominant HAB concern in New England coastal waters. This has begun to change with the emergence of amnesic shellfish poisoning (ASP) and diarrhetic shellfish poisoning (DSP) syndromes, caused by blooms of Pseudo-nitzschia spp. and Dinophysis spp., respectively. Other fin- and shellfish killing species are also increasingly impacting aquaculture and fisheries in the region. This project addresses these challenges through development of HABON-NE (HAB Observing Network - New England), a framework for aligned academic, industry, state, and federal scientists to deploy a fleet of advanced sensors and sensor platforms. Data and analytical products are shared as they are created through the WHOI HAB Hub, an open source webserver platform for collating rich and diversely sourced HAB observations, model outputs, and management actions. Check out HAB Hub here.
Co-PIs: Collin Roesler (Bowdoin College); Kate Hubbard (FWRI); Jake Kritzer (NERACOOS); Greg Doucette, Yizhen Li, Rick Stumpf (NOAA); Donald Anderson, Mindy Richlen, and Bruce Keafer (WHOI)
Funding: NOAA NCCOS MERHAB program
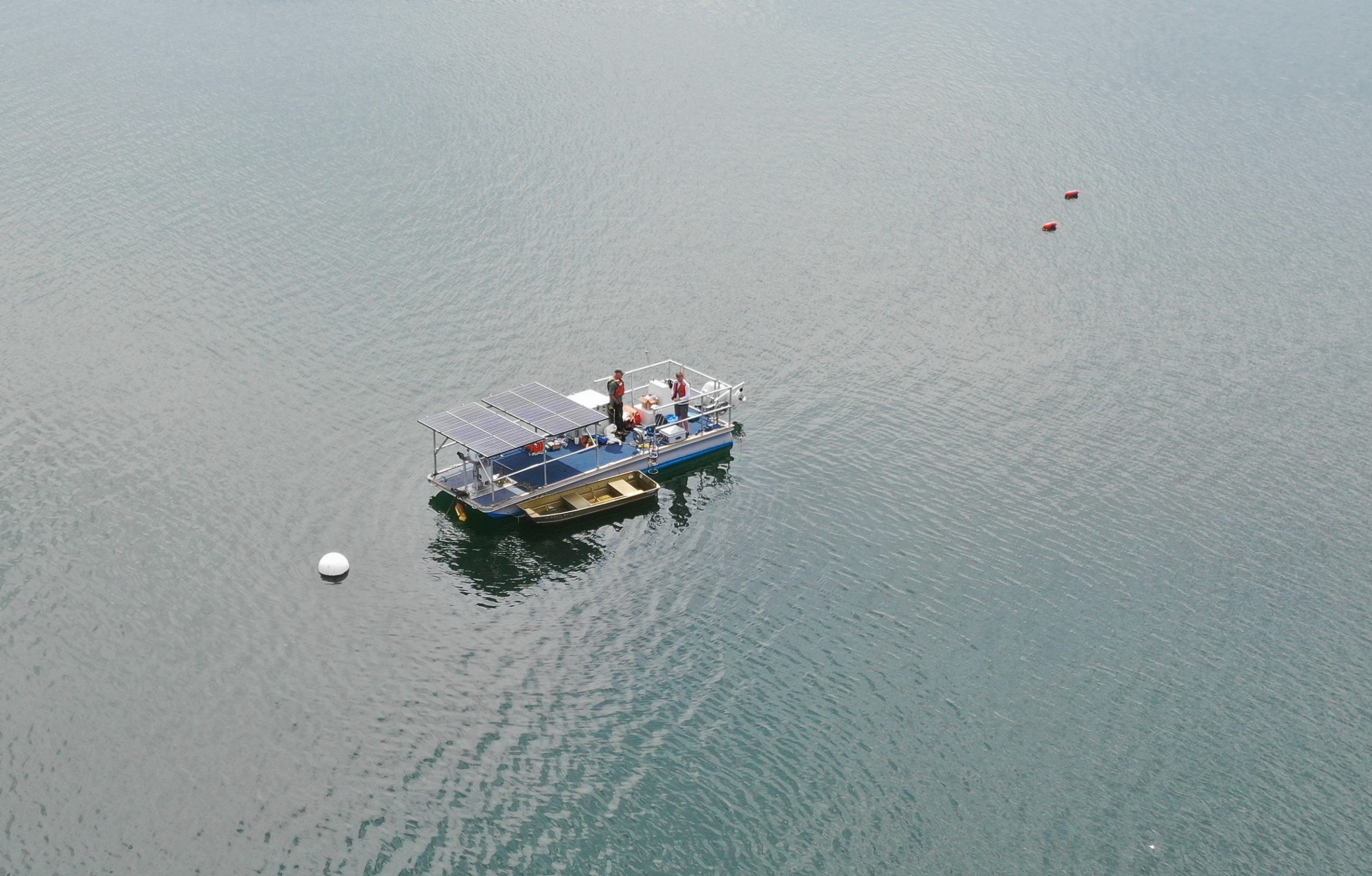

PhytO-ARM, an open-source platform for real-time phytoplankton monitoring, data sharing, and automated aquaculture management
PhytO-ARM (Phytoplankton Observing for Automated Real-time Management) is a ROS-based, open-source platform for multi-sensor real-time phytoplankton observing. Our goal is to enable description of the diversity, abundance, and vertical distribution of phytoplankton through the water column. Sensors are integrated and controlled through software developed for the Imaging FlowCytobot (IFCB) sensor. Vertical profiling and sensor positioning is made possible through low-cost, networked winches that are built from commercial-off-the-shelf components. Additional capabilities are being developed so that PhytO-ARM systems can detect HABs in near real-time and take actions to protect HAB-sensitive animals at aquaculture facilities.
Co-PIs: Donald Anderson (WHOI), Kate Hubbard, Célia Villac (FWRI), and Greg Doucette (NOAA)
WHOI contributors: Verena Lucke, Ryan Govostes, Sidney Batchelder, and Bob Petitt
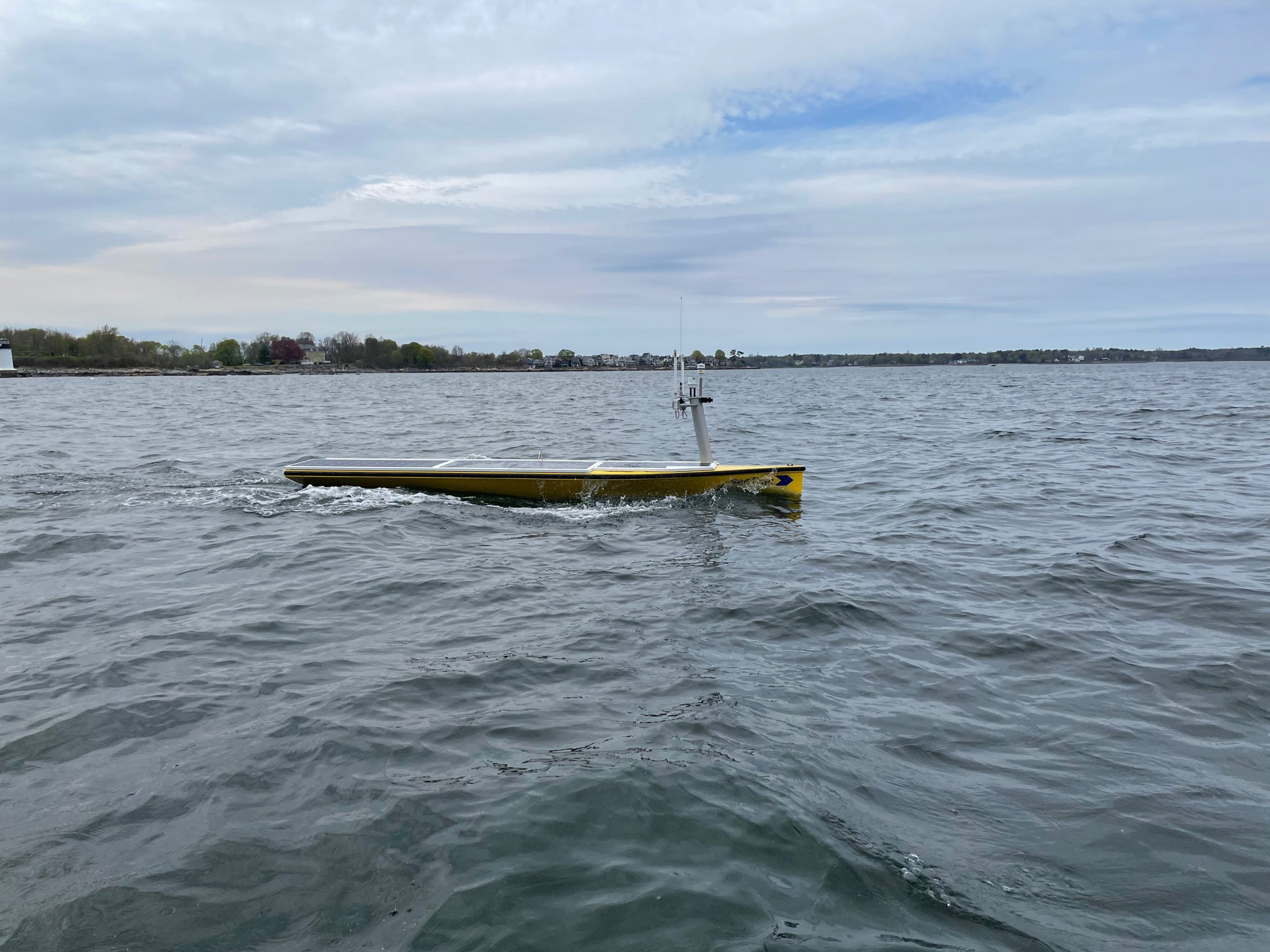

Persistent Autonomous Mobile Monitoring of Waterborne Biochemical Agents
This project aims to demonstrate an autonomous surface vehicle (ASV) made by SeaTrac Systems as a deployment platform for a range of chemical and biological sensors. The WHOI team's focus is on integration of the IFCB sensor and testing of the boat in marine settings. We are working with McLane Research Labs to update the IFCB for use aboard the SeaTrac ASV and using our PhytO-ARM platform for collection of underway data by other sensors.
Lead-PI: Robert Moorhead (Mississippi State University); Other Co-PIs: Daniel Chesser, Wes Lowe, and Dash Padmanava (MSU)
WHOI contributors: Cameron Fairclough
Funding: Army ERDC
Investigation of Type II Syndiniales-host interactions through single cell transcriptomics
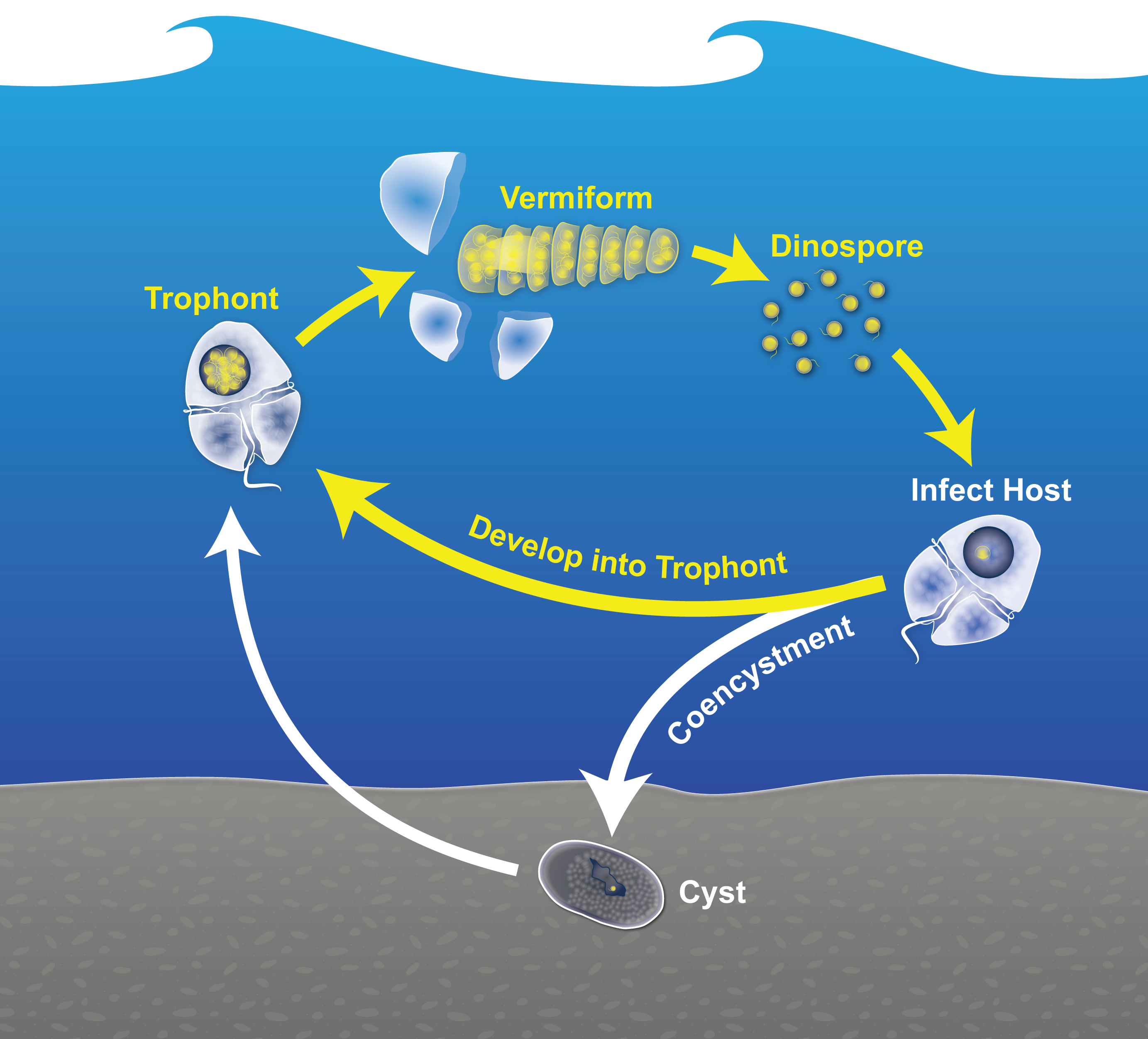
Parasitic protists play a central role in marine food webs as top-down controls on a wide range of protist and animal prey. Recent global surveys have highlighted the dominance of Group II Syndiniales parasites across diverse marine environments. This project focuses on one of these species, Amoebophrya ceratii, and its interactions with its host, the HAB dinoflagellate Alexandrium catenella. During blooms, A. ceratii invades the cytoplasm of A. catenella then proliferates to produce hundreds of new infective dinospores. When new hosts become scare it may instead co-encyst with A. catenella, using its hosts' cyst stage as a refugium between blooms. We are exploring interactions between these species to determine the molecular signals that may drive co-encystment or further parasite growth. To do this, we are combining new lab- and field-based approaches including adaptive thin layer sampling and a high throughput approach for collection of single cell transcriptomes. The latter is based on SeqWell, a method developed by the team in Co-PI Alex Shalek's lab.
Lead-PI: Harriet Alexander (WHOI); Other Co-PIs: Ginny Edgcomb, Maria Pachiadaki (WHOI); Alex Shalek (MIT)
Funding: Gordon and Betty Moore Foundation - Symbiosis in Aquatic Systems
